Subcortical segmentation package¶
This package contains a deep learning model for segmenting four subcortical structures: the cerebellum (CB), lateral posterior ventricle horns (LPVH), choroid plexus (ChP), and the cavum septum pellucidum et vergae (CSPV).
The model expects images aligned to the atlas coordinate system as input, that are in their original size (i.e. no scaling parameters applied). Segmentations for aligned scaled images can be obtained by applying the scaling factor to the segmentations. Segmentations for images in their original alignment can be obtained by applying the inverse alignment transform to the segmentations. Examples on these workflows are provided in the example gallery.
An example subcortical segmentation can be produced with the following code:
from fetalbrain.structural_segmentation.subcortical_segm import segment_scan_subc
from fetalbrain.utils import read_image, plot_planes_segm
from fetalbrain.alignment.align import align_scan
from fetalbrain.model_paths import EXAMPLE_IMAGE_PATH
# load an image
example_scan, _ = read_image(EXAMPLE_IMAGE_PATH)
# align the scan to the atlas space without scaling
aligned_scan, params = align_scan(example_scan, scale=False)
# perform segmentation on aligned image
segmentation, structure_names = segment_scan_subc(aligned_scan, connected_component=True)
# plot the segmentation
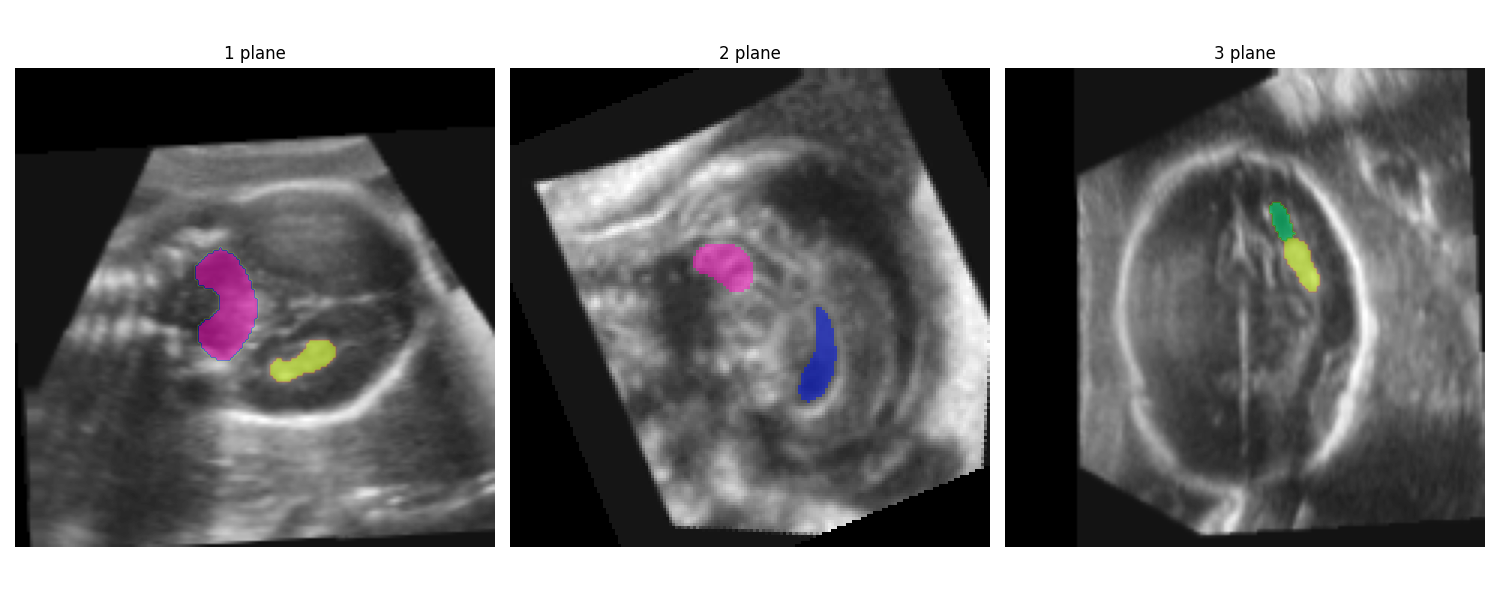
fig = plot_planes_segm(aligned_scan.squeeze(), segmentation.squeeze())
Running this results in the following output figure (CB: pink, ChP: yellow, LPVH: green, CSPV: blue):
More details about the segmentation functions are provided in the
Credits¶
The code in this package has mainly been developed by Linde Hesse, and the methods are described in detail in a NeuroImage paper. Please cite this paper if you use the subcortical segmentation model in your research:
The trained network weights included in the package are from a model trained on a subset of 20 manually annotated fetal brain volumes (rather than only 9 as described in the paper).