Normative Growth Trajectories¶
Introduction¶
Here we provide guidance on how to use this repo to generate the results seen in: Normative growth trajectories of fetal brain regions validated by satisfactory maturation of neurodevelopmental domains at 2 years of age
Installation¶
Install the code following steps in Getting started
Pipeline Example¶
The following example implements the pipeline used in Normative Growth Trajectories manuscript.
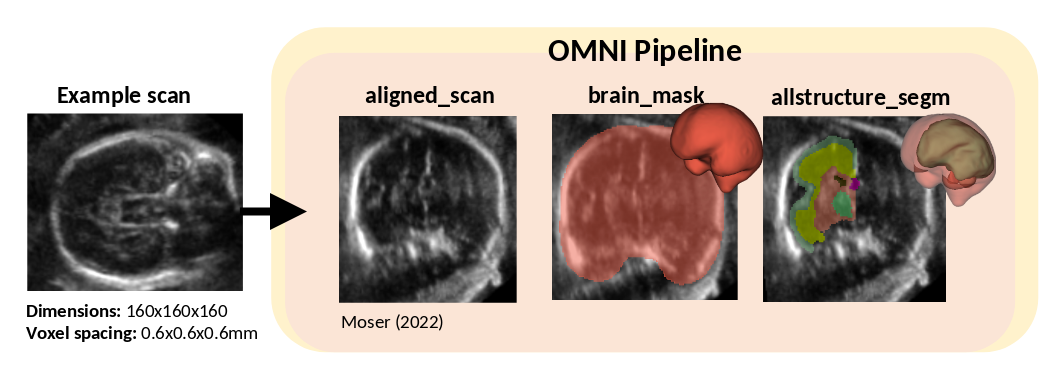
This pipeline will save out, an aligned scan aligned_scan.nii.gz, a brain mask used to compute TBV brain_mask.nii.gz and the 15 region segmentation allstructure_segm.nii.gz. Segmentation index:
“CoP”: 1, “CSP”: 2, “CB”: 3, “ChP”: 4, “LV”: 5, “DGM”: 6, “Th”: 7, “BS”: 8, “WM”: 9, “FH”: 10,
To run the pipeline for multiple scans and with more flexibility, it is recommended to use the individual pipeline functions rather than the wrapper functions. This ensures that the models are not reloaded for each scan. The following example demonstrates this for a single example.
To ammend for your data: change: $EXAMPLE_IMAGE_PATH and $savefolder
from pathlib import Path
import torch
from fetalbrain.utils import read_image, write_image
from fetalbrain.alignment.align import load_alignment_model, align_to_atlas, prepare_scan
from fetalbrain.tedsnet_multi.teds_multi_segm import (
load_tedsmulti_model,
segment_tedsall,
load_sidedetector_model,
detect_side,
)
from fetalbrain.brain_extraction.extract import extract_brain, load_brainextraction_model
from fetalbrain.model_paths import EXAMPLE_IMAGE_PATH
# Load the models once
align_model = load_alignment_model()
teds_multimodel = load_tedsmulti_model()
side_detectormodel = load_sidedetector_model()
brainextraction_model = load_brainextraction_model()
# ------------------------------------------------
# Loop over all scans (just one here as example)
# Here you can change EXAMPLE_IMAGE_PATH to your
# image path and/or loop through your directory
# for volumes!
example_scan, _ = read_image(EXAMPLE_IMAGE_PATH)
# ------------------------------------------------
torch_scan = prepare_scan(example_scan)
# Start with alignment to atlas space
aligned_scan, params = align_to_atlas(torch_scan, align_model, scale=False)
# Perform segmentation with multi structure tedsnet
side, prob_side = detect_side(aligned_scan, side_detectormodel)
allstructure_segm, multi_keys = segment_tedsall(aligned_scan, teds_multimodel, side=side)
# perform whole brian extraction (i.e. brain masking)
brain_mask, brain_key = extract_brain(aligned_scan, brainextraction_model)
# ------------------------------------------------
# Write out the results in the aligned orientation
# you can change this to where you want your results saved!
savefolder = Path("results")
# ------------------------------------------------
savefolder.mkdir(exist_ok=True)
write_image(savefolder / "aligned_scan.nii.gz", aligned_scan.squeeze().numpy())
write_image(savefolder / "allstructure_segm.nii.gz", allstructure_segm.squeeze(), segm=True)
write_image(savefolder / "brain_mask.nii.gz", brain_mask.squeeze().numpy(), segm=True)
Compute Volume¶
To compute the volume of each segmented structure, this minimal example can be used:
import numpy as np
from pathlib import Path
from fetalbrain.utils import read_image
def ComputeVol(allstructure_segm, brain_mask,voxel_dim=0.6,N=160):
assert np.shape(allstructure_segm) ==np.shape(np.zeros((N,N,N)))
assert np.shape(brain_mask) ==np.shape(np.zeros((N,N,N)))
struc_vol_dict = {}
# compute the TBV:
volume = np.count_nonzero(brain_mask)*(voxel_dim**3)
struc_vol_dict["TBV"] =volume
# Index of each of the structures
key_maps = {
"CoP": 1,
"CSP": 2,
"CB": 3,
"ChP": 4,
"LV": 5,
"DGM": 6,
"Th": 7,
"BS": 8,
"WM": 9,
"FH": 10,
}
for keys in key_maps.keys():
""" Loop through the structures and measure volume
"""
struc = np.where(allstructure_segm==key_maps[keys],1,0)
volume = np.count_nonzero(struc)*(voxel_dim**3)
struc_vol_dict[keys+"V"] =volume
print(struc_vol_dict)
if __name__ == '__main__':
""" Compute the volume measures of the segmented structures
"""
savefolder = Path("results")
allstructure_segm,h = read_image(savefolder / "allstructure_segm.nii.gz")
brain_mask,h = read_image(savefolder / "brain_mask.nii.gz")
# Measure the volume of each structire:
SVol = ComputeVol(allstructure_segm, brain_mask)
Compare Volume¶
The following example shows how to compare your volume measure to the normative growth equations (Suppl Table 8)
import math
import numpy as np
class NormGrowthEQ:
def __init__(self) -> None:
pass
"""
Normative Growth Trajectories for volume in cm3
Input: GA - the gestational age of the subject (in weeks)
"""
def TBV(self,GA):
u = 1.954510 + 0.018205*GA**3 + -0.178633*GA**2
o = np.exp(-0.702623 + 0.150265*GA)
return u,o
def CoPV(self,GA):
u = 213.602100 + -1055185.000000*GA**-3 + 179286.700000*GA**-2 + -10465.890000*GA**-1
o = np.exp(-3.319410 + 0.276768*GA + -0.004899*GA**2)
return u,o
def WMV(self,GA):
u = 147.933791 + -20.171007*GA + 0.862044*GA**2 + -0.010213*GA**3
o = np.exp(8.181534 + -34.670966*GA**-0.5)
return u,o
def DGMV(self,GA):
u = 2.565062 + -0.905458*GA**0.5 + 0.000642*GA**3
o = np.exp(-3.357278 + 0.134731*GA)
return u,o
def CBV(self,GA):
u = 20.062157 + -26.210291*math.log(GA) + 13.249654*GA**0.5
o = np.exp(-4.802530 + 0.144062*GA)
return u,o
def ChPV(self,GA):
u = -8.747354 + 1.226594*GA + -0.052478*GA**2 + 0.000745*GA**3
o = np.exp(-4.017505 + -1353.340000*GA**-2 + 110.794500*GA**-1)
return u,o
def LVV(self,GA):
u = 0.199152 + -759.201300*GA**-3 + 0.000006*GA**3
o = np.exp(-8.870769 + 1.868799*math.log(GA))
return u,o
def FHV(self,GA):
u = 1.957187 + -0.247137*GA + 0.010655*GA**2 + -0.000135*GA**3
o = np.exp(-28.094510 + 1841.363000*GA**-1 + 334027.300000*GA**-3 + -43795.430000*GA**-2)
return u,o
def BSV(self,GA):
u = 27.563862 + -187.829409*GA**-0.5 + 9843.864509*GA**-2 + -77842.728897*GA**-3
o = np.exp(41.292163 + -7.207325*math.log(GA) + -745.053409*GA**-1 + 5815.331813*GA**-2)
return u,o
def ThV(self,GA):
u = -0.006779 + 0.000063*GA**3
o = np.exp(2.222630 + -21.555120*GA**-0.5)
return u,o
def CSPV(self,GA):
u = 4.356401 + -0.607443*GA + 0.026988*GA**2 + -0.000357*GA**3
o = np.exp(-7.132420 + 0.186246*GA)
return u,o
if __name__ == "__main__":
# Set up the class containing all equations
EQ = NormGrowthEQ()
# Individual
CoP_volume = 7 #cm3
age = 20.2 #gestational weeks
# Compute z-score
u,o = getattr(EQ,"CoPV")(age) # find normative mean + std
z = (CoP_volume - u)/o
print("z-score for individual: ", z)